anvio
if you successfully installed anvio, you can proceed from here.
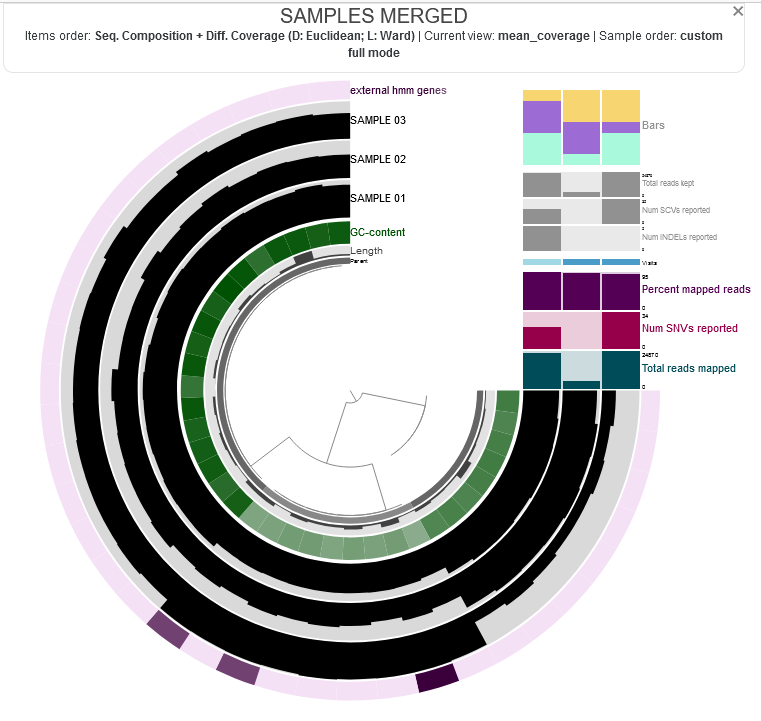
anvio visualizations look like this:
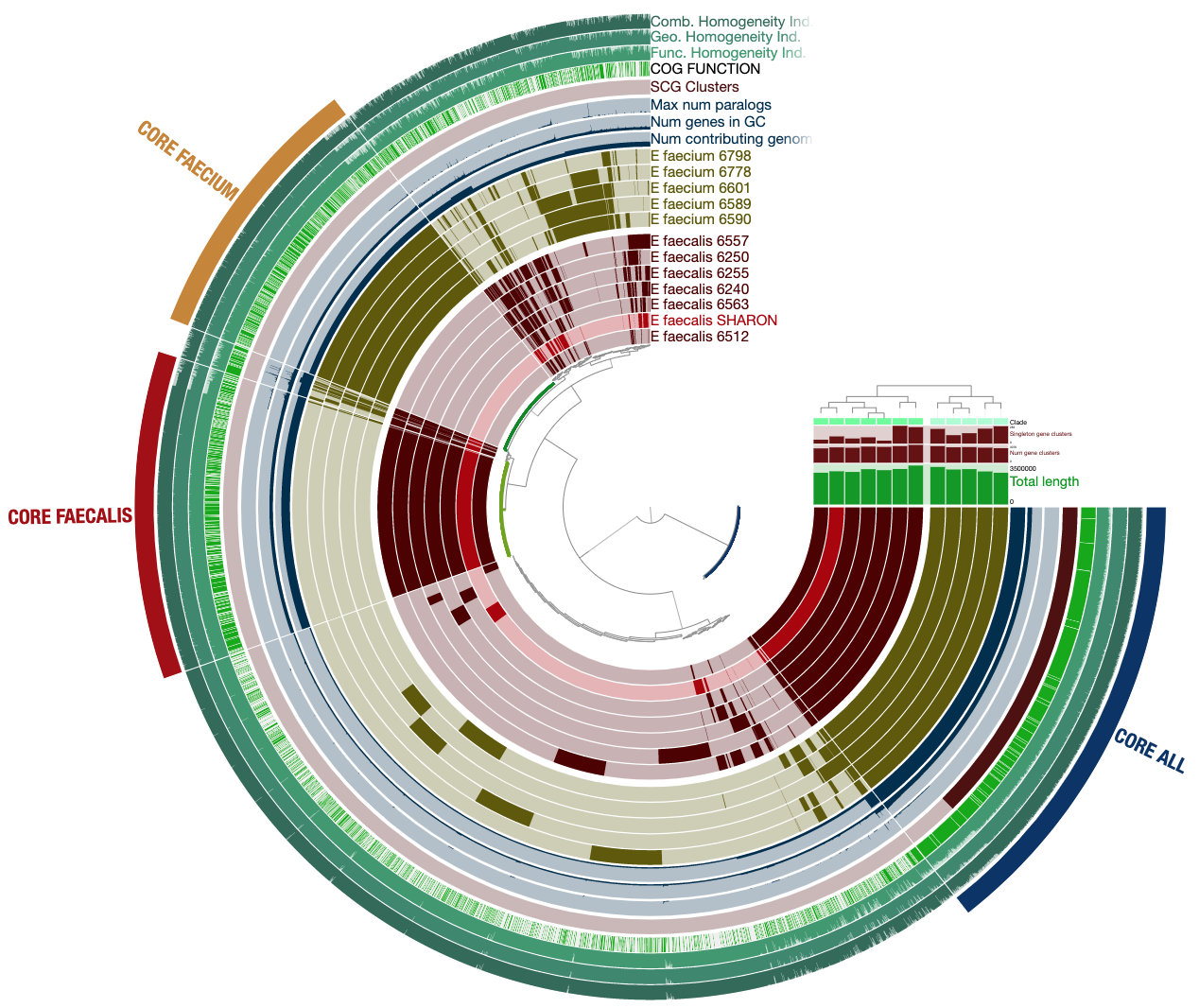
when you learn to generate it, you can interact with it by hovering your mouse over each part to see more information.
now we need to check if this interactive command works correctly, even though we don’t have any data or files to visualize yet. we actually need some files with the .db extension, but that’s okay—we’ll fake some. we’ll enter this code and expect to get an error saying the files don’t exist, which confirms that anvi-interactive works perfectly. however, if we get another message related to the interactive part, we’ll fix that.
anvi-interactive -c CONTIGS.db -p MERGED/PROFILE.db -I localhost
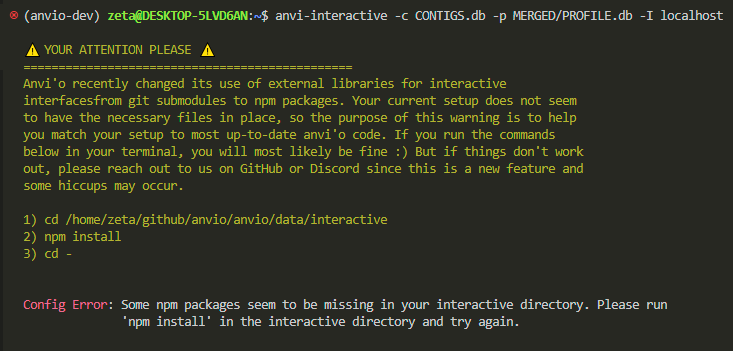
i got the nicest error message, explaining in plain language how to fix the problem. so, i need to set my path correctly, install something called npm, and then run cd -.
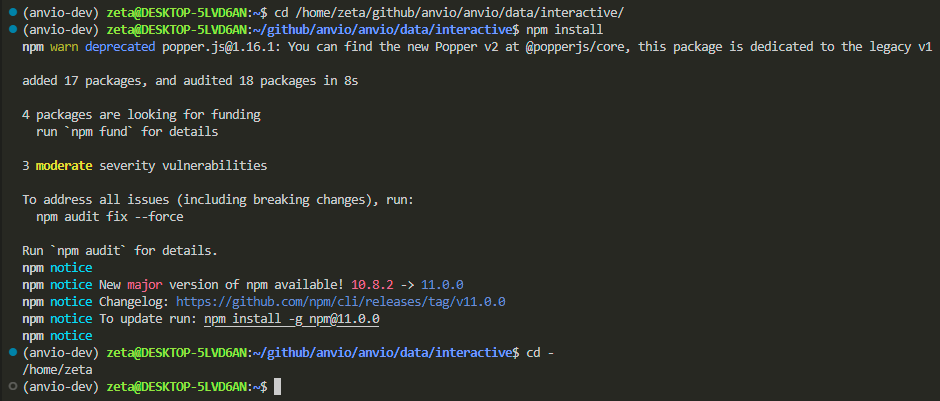
let’s run anvio interactive with our fake files again:
anvi-interactive -c CONTIGS.db -p MERGED/PROFILE.db -I localhost
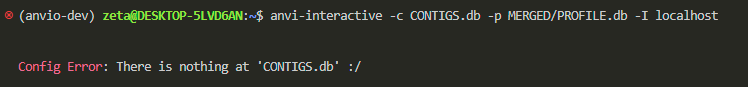
now we got the correct error saying there’s no data in your system yet.
now, let’s ask anvio to do a full self-test:
anvi-self-test --suite mini
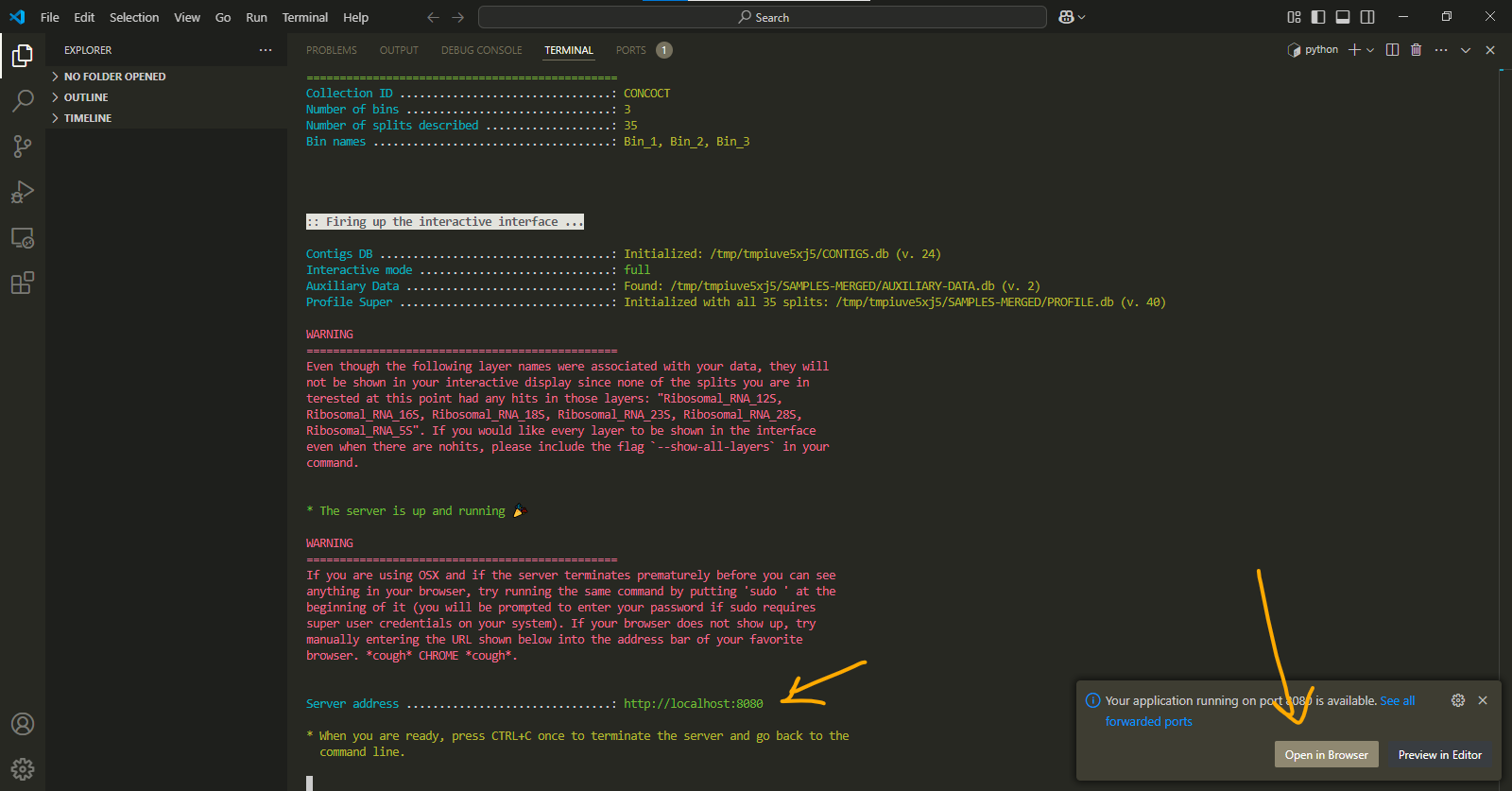
this test will show us how a real anvi-interactive works. by opening your localhost in the browser, you can see the newest version and a visualization.
localhost is a url generated on your system to view visualizations. it can have either of these addresses:
http://localhost:8080/
http://127.0.0.1:8080/
anvio visualizations work best with chrome. so, if you notice any weirdness, try opening the url in chrome.
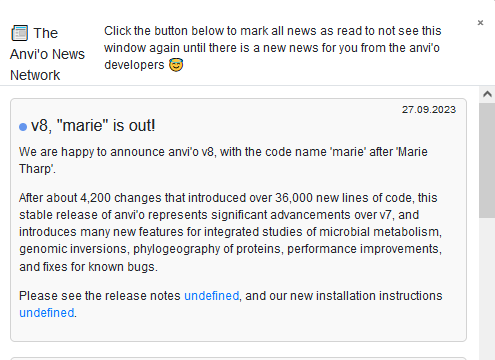
also, kudos to whoever is naming the versions. 🎉
finalize and prepare for everything
we know that not all parts of a genome are translated into proteins. there are specialized tools that help identify the functional regions of a genome and tag them with their taxonomical or functional roles. anvio allows us to integrate these tools and databases.
let’s go:
anvi-setup-scg-taxonomy
it took me less than a minute with the university internet nework, it might vary for you.
(i’ll mention the time it took me for a relative comparison.)
anvi-setup-ncbi-cogs
6 minutes
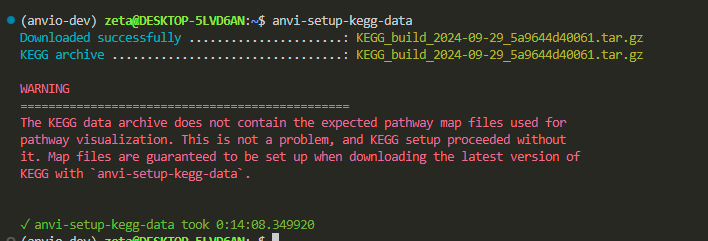
anvi-setup-kegg-data
14 minutes
now do another test:
anvi-self-test --suite pangenomics
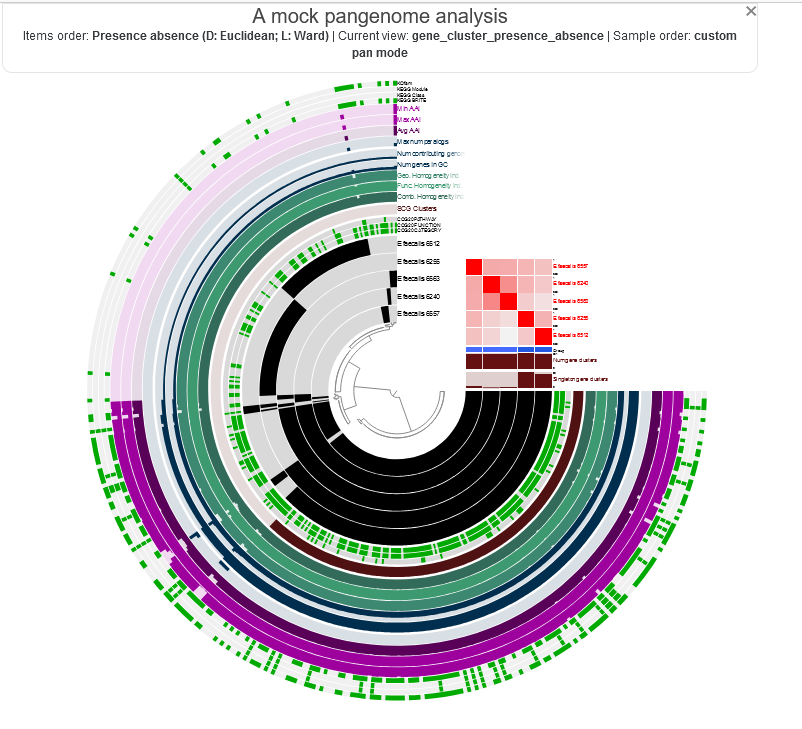
i seem to be safe. how is it going for you over there? :)
binning algorithm
you will learn what binning means during the course, but for now, we’ll just set up the necessary tools for it.
move to the github directory:
cd ~/github/
clone CONCOT to anvio conda enviornment:
git clone https://github.com/merenlab/CONCOCT.git
cd CONCOCT
python setup.py build
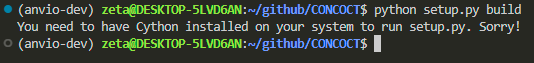
i need cython. i don’t know what it is either, but let’s go ahead and figure out how to install it anyway.
conda install -c conda-forge cython
cython --version
re-run:
python setup.py build
python setup.py install
finished! now we run this command to see a page like this:
anvi-cluster-contigs -h
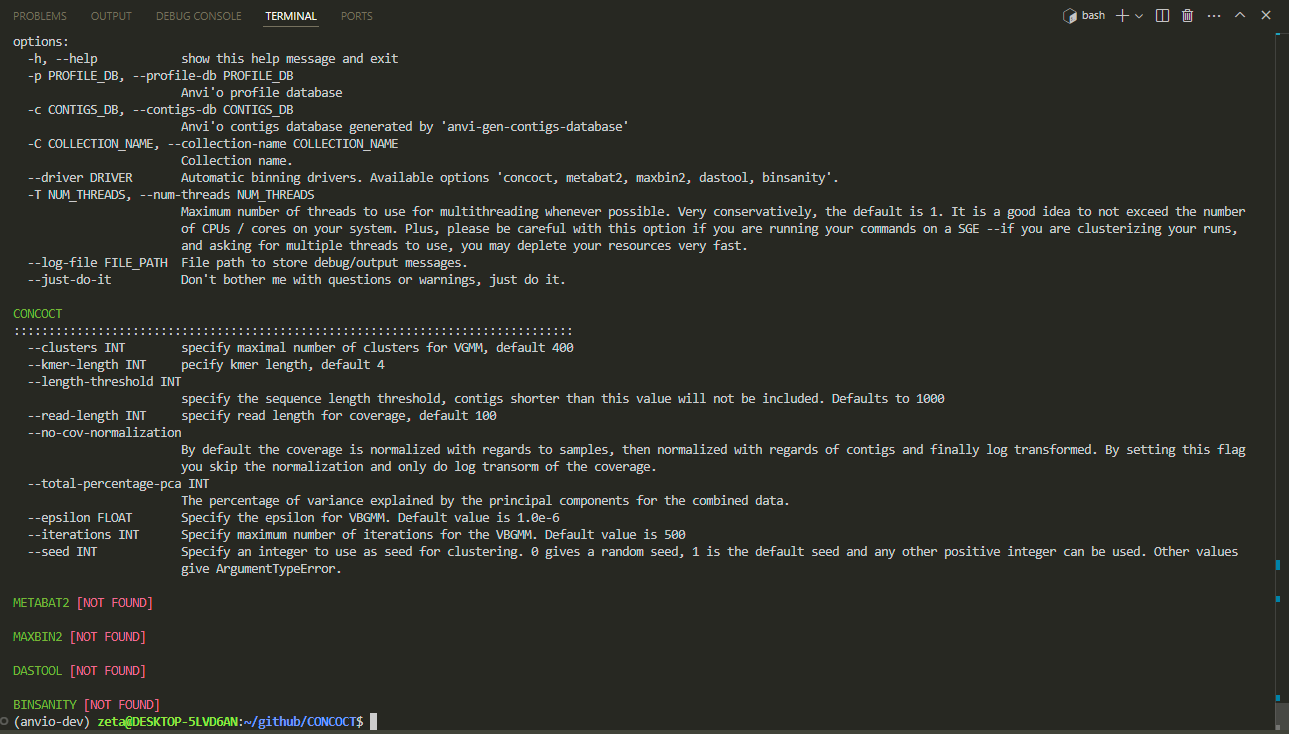
we can go for a cup of tea now :D